A Genetic Kundli is a genomics-based health platform that decodes your DNA to reveal both the health risks you were born with and the genetic changes acquired throughout your life.
Genetic variations present from birth, shared across all cells and passed through generations.
Genetic changes that develop during life, associated with diseases such as cancer and not inherited.
Discovery of the BRCA1 gene demonstrated how inherited mutations significantly increase the risk of breast and ovarian cancer.
Miki Y et al., 1994
View Publication →Sequencing the human genome established the foundation for understanding how genes influence health, disease risk, and biological traits.
Lander ES et al., 2001
View Publication →GWAS research identifies genetic variants influencing diseases such as diabetes, cardiovascular disease, and autoimmune disorders.
McCarthy MI et al., 2008
View Publication →Research shows that genetic variants contribute to inherited susceptibility for many complex diseases.
Manolio TA et al., 2009
View Publication →Precision medicine integrates genomic information with clinical data to guide personalized healthcare strategies.
Ashley EA, 2016
View Publication →Genome-wide association studies have identified thousands of genetic variants linked to complex human diseases and traits.
Visscher PM et al., 2017
View Publication →From Sample to Scientific Insight - How We Build Your Personalized Genetic Kundli
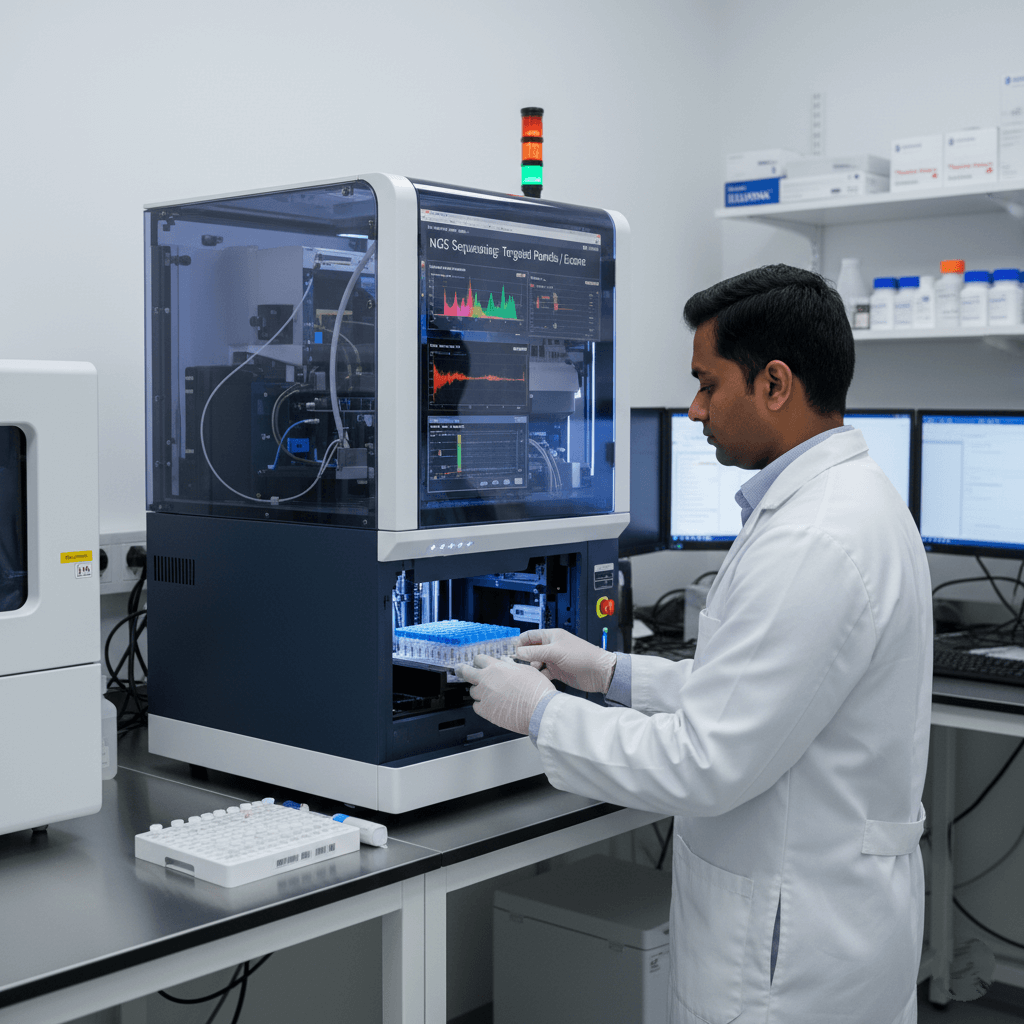

Sequencing means reading the exact order of the DNA letters (A, T, C, G) in your sample, just like reading a book, one word at a time.
QC = Quality Checks. It’s like a double-check system at every step to make sure the DNA reading is accurate, complete, and trustworthy before sending results to your doctor.
| Step | What Happens | Why It Matters |
|---|---|---|
| 1. DNA Preparation | Your sample arrives. DNA is extracted and broken into tiny pieces. | Clean DNA = accurate reading. |
| 2. Library Preparation | Tags (adapters) added so machines can read DNA. | Allows millions of pieces to be read at once. |
| 3. Target Capture | Magnetic beads grab only key genes (e.g., BRCA1, TP53). | Saves time — skips junk DNA. |
| 4. Sequencing | DNA flows in a flow cell. Camera reads colored lights (A=green, etc.). | Generates billions of 150-letter reads. |
| 5. QC Checks | Computers check clarity, balance, accuracy. | Catches errors early. |
| Metric | Target | What It Means |
|---|---|---|
| Uniformity | ≥90% | All genes read evenly — no missed spots. |
| On-Target Rate | ≥70% | Most reads are from intended genes. |
| Coverage Depth | 100x–1000x | Each letter read 100–1000 times. |
| Error Rate | <1 in 10,000 | Extremely low error chance. |
Example: If a gene has a mutation, we need to see it clearly and repeatedly — not just once by chance.
| Type | What It Covers | Best For |
|---|---|---|
| Targeted Gene Panel | 50–500 cancer genes | Fast, affordable, focused |
| Whole Exome | ~22,000 protein-coding genes | Broader screening |
| Hybrid-Capture / Amplicon | Two DNA capture methods | Both highly accurate |
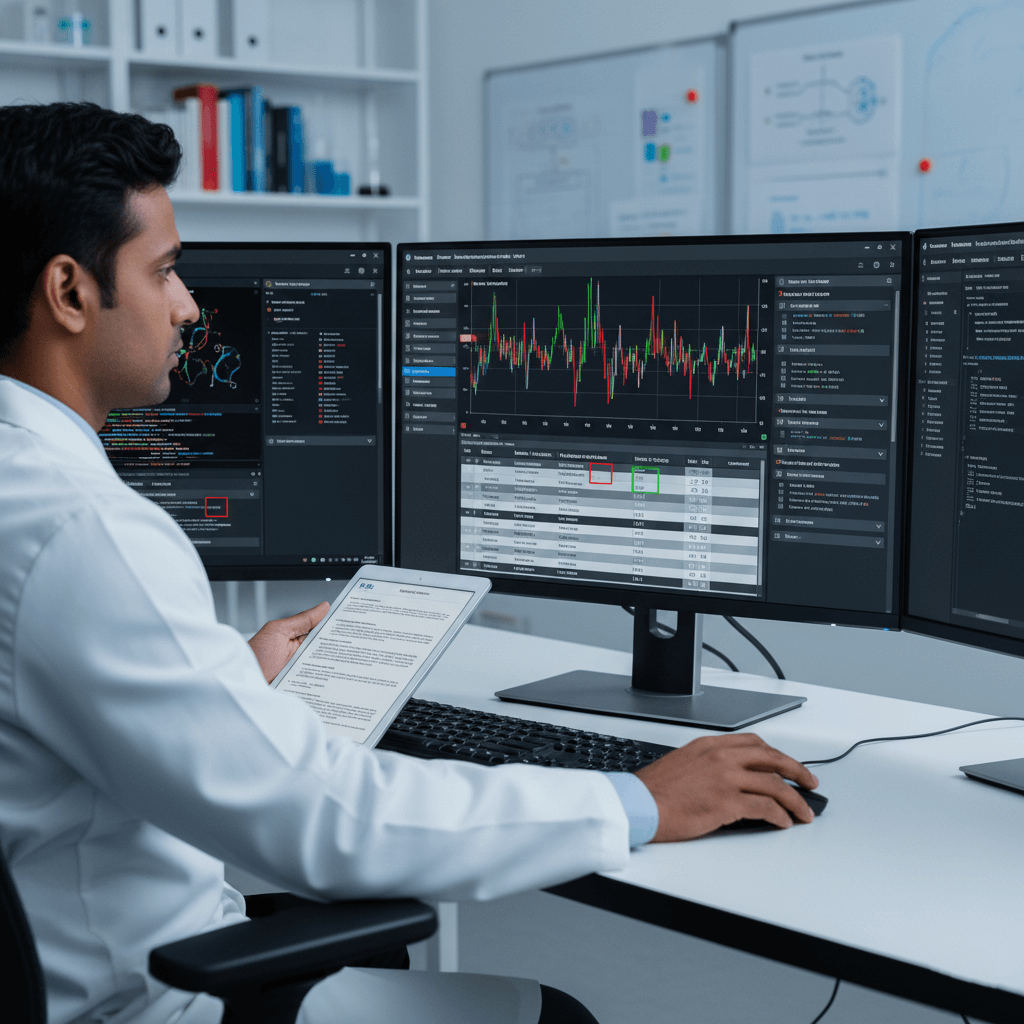

Bioinformatics = Turning raw DNA data into meaningful health insights.
Think of it like this: Step 2 gave us billions of DNA “words” Step 3 reads them, finds spelling mistakes (mutations), and tells us what they mean for your health.
We use smart computers + expert scientists to do this accurately and fast.
Our BWA/GATK-class pipeline is the gold standard in clinical genomics, used by top labs worldwide. It’s automated but never fully automatic every result is reviewed by a human expert.
| Step | What Happens | Why It Matters |
|---|---|---|
| 1. Align DNA Reads | Billions of short DNA pieces are lined up to the human genome map using BWA (like putting puzzle pieces in place). | Ensures we know exactly where each DNA letter belongs. |
| 2. Clean & Refine | Fix errors from the machine (e.g., duplicates, misreads) using GATK tools. | Removes noise so only real changes remain. |
| 3. Find Variants | Look for spelling mistakes (SNPs, indels), missing chunks (CNVs), and rearrangements (SVs). | Catches all types of DNA changes, not just small ones. |
| 4. Name Changes Correctly | Use HGVS nomenclature the universal language of genetics (e.g., c.123A>G). |
Doctors worldwide understand the exact mutation. |
| 5. Pick the Right Transcript | Genes have multiple “versions” we pick the most trusted one (e.g., MANE) using best practices. | Avoids confusion same gene, same meaning. |
| 6. Add Meaning (Annotation) | Compare variants to ClinVar (known diseases) and gnomAD (normal population). | Tells us: “Is this dangerous, harmless, or unknown?” |
| Feature | What It Means for You |
|---|---|
| BWA/GATK-class | Same tech as Harvard, Broad Institute, NHS clinically proven. |
| CNV/SV Support | Detects large deletions, duplications, inversions critical in cancer. |
| HGVS Nomenclature | Your mutation is written in standard medical code no ambiguity. |
| Best-Practice Transcripts | We use MANE/RefSeq the most reliable gene versions. |
| ClinVar + gnomAD | Backed by millions of real patient & healthy DNA samples. |
| Expert Review | A PhD bioinformatician + geneticist checks every report. |
Example: From Data to Diagnosis
A instead of G at position 17,234,567 →
HGVS: c.35G>A in BRCA1 →
Pathogenic (per ClinVar) → High breast cancer risk
| Type | Example | Health Impact |
|---|---|---|
| SNV | Single letter change | Can turn a gene on/off |
| Indel | Small insert/delete | May break a protein |
| CNV | Whole gene duplicated/deleted | Often linked to cancer |
| SV | Large rearrangement | Can fuse two genes |

Reporting = Turning complex DNA results into a clear, actionable health report your Genetic Kundli.
It’s the final step where science meets care.
| Feature | What It Means for You |
|---|---|
| Tiered Variants | Ranked by importance: Tier 1 (high risk) to Tier 4 (benign) |
| Actionability Notes | Clear next steps (e.g., “start screening”) |
| Therapy Pointers | Personalized drugs (e.g., PARP inhibitors) |
| Clinical Trials | Eligible studies near you (informational) |
| Re-analysis | Free update if science changes meaning |
| What It Is | Why It Matters |
|---|---|
| Pre-Test Counseling | Understand risks, benefits, family impact |
| Post-Test Counseling | Explain results, answer fears |
| Family Planning | Implications for children/siblings |
| Emotional Support | Process anxiety, guilt, or relief |
| Follow-Up Plan | Schedule screenings or referrals |
Included Free: 1-hour video call with a certified genetic counselor (CGC).
Example: From Variant to Action
BRCA1 pathogenic → Tier 1 – High Risk → Mammogram every 6 months + consider preventive options
Indicative catalogue with enhanced gene coverage & variant detection (2025 Edition). Panels evolve with NCCN, ELN, ACMG, CPIC guidelines. Custom panels available.
Germline • Inherited Risk Panels
Germline risk assessment panels for inherited cancer syndromes. Includes ACMG SF v3.2 (81 genes).
Full intron coverage and secondary findings reporting.
| Panel | Key Genes |
|---|---|
| Breast/Ovarian | BRCA1/2, PALB2, CHEK2, ATM, PTEN, TP53, RAD51C/D, BRIP1, ACMG SF v3.2 NEW |
| Lynch Syndrome | MLH1, MSH2, MSH6, PMS2, EPCAM |
| Polyposis | APC, MUTYH (biallelic), STK11, SMAD4, BMPR1A, POLE/POLD1 |
| Endocrine | RET (MEN2), MEN1, VHL, SDHx (SDHB, SDHC, SDHD) |
| Li-Fraumeni | TP53 comprehensive, ACMG SF v3.2 NEW |
Covers polygenic risk for T2D and rare monogenic forms; essential & monogenic hypertension.
| Panel | Key Genes |
|---|---|
| T1D | HLA-DRB1/DQB1, PTPN22, CTLA4 |
| T2D | TCF7L2, FTO, KCNJ11, ABCC8, PPARG, PRS NEW |
| MODY | GCK, HNF1A, HNF4A, HNF1B |
| Neonatal | KCNJ11, ABCC8, INS |
| Essential Hypertension | ACE, AGT, AGTR1, NOS3 |
| Liddle | SCNN1A/B/G |
Inherited cardiomyopathy, arrhythmia, and aortopathy. Guides ICD placement and family screening.
| Panel | Key Genes |
|---|---|
| HCM/DCM/ARVC | MYH7, MYBPC3, TTN, DSP, PKP2, LMNA, TNNI3, TNNT2, FLNC, TTR NEW |
| Channelopathies | KCNQ1, KCNH2, SCN5A (LQTS, Brugada) |
| Aortopathy | FBN1 (Marfan), TGFBR1/2, SMAD3, ACTA2, DSG2, TMEM43 NEW |
Epilepsy, neuromuscular, ataxia/leukodystrophy, and selected neurodegenerative diseases.
| Panel | Key Genes |
|---|---|
| Epilepsy | SCN1A, SCN2A, KCNQ2/3, CDKL5, PCDH19, STXBP1, KCNT1, SLC2A1, PRRT2 NEW |
| Neuromuscular | DMD, SMN1/2 (CNV), RYR1, COL6A1-3, GAA repeat (FXN) NEW |
| Ataxia/Leukodystrophy | FXN (FRDA), ATXN1/2/3, ABCD1, EIF2B |
| Neurodegeneration | APP, PSEN1/2, MAPT, GRN, LRRK2, GBA |
Carrier screening, invasive prenatal diagnostics, and NIPT (screening only).
| Panel | Key Features |
|---|---|
| Expanded Carrier | CFTR, HBB/HBA, SMN1, GJB2, PAH, G6PD (100–500 genes), FMR1 repeats NEW |
| Prenatal Diagnostic | Aneuploidy + microdeletions (CMA), single-gene, 22q11, 1p36 NEW |
| NIPT* | Trisomy 21/18/13; sex chromosomes, microdeletion panel NEW |
For undiagnosed developmental delay, metabolic, and mitochondrial disorders.
| Panel | Key Features |
|---|---|
| Clinical Exome | SNV/indel + CNV (trio), annual re-analysis NEW |
| Genome | SVs, non-coding, mtDNA, regulatory variants NEW |
| Mitochondrial | Full mtDNA + heteroplasmy |
Medication response & safety. CPIC-aligned reporting.
| Gene | Clinical Use |
|---|---|
| CYP2D6/2C19/2C9, VKORC1 | Psych, PPI, anticoagulants |
| SLCO1B1 | Statin myopathy |
| DPYD, UGT1A1 | Oncology dosing |
| HLA-B*15:02/*57:01 | Carbamazepine, abacavir |
| TPMT, NUDT15 | Thiopurines NEW |
| CYP3A5 | Tacrolimus NEW |
| IFNL3/4 | Hepatitis C NEW |
Somatic • Tumor-specific Panels
Site-specific and pan-cancer options for therapy selection & hereditary risk. Detects actionable SNVs/indels, CNVs, fusions, TMB, MSI.
Enhanced with deep intronic coverage & RNA fusion analysis. NCCN/ESMO 2025 compliant.
| Panel | Key Genes / Biomarkers |
|---|---|
| Breast/Ovarian | BRCA1, BRCA2, PALB2, CHEK2, ATM, PTEN, TP53, CDH1, RAD51C/D, BRIP1 NEW |
| Lung (NSCLC) | EGFR, ALK, ROS1, BRAF, KRAS (G12C), MET exon14, RET, NTRK1-3, HER2/ERBB2, TMB ≥10 mut/Mb NEW |
| Colorectal | KRAS/NRAS, BRAF, MSI/dMMR (MLH1, MSH2, MSH6, PMS2), APC, PIK3CA, HER2 (CNV) NEW |
| Prostate | BRCA1/2, ATM, CHEK2, HOXB13, PTEN, MSH2, CDK12, MSI NEW |
| Melanoma | BRAF, NRAS, KIT, NF1 |
| Glioma | IDH1/2, 1p/19q (CNV), TERT, ATRX, TP53 |
| Pan-Cancer | 50–500 genes: SNVs/indels, CNVs, fusions, TMB, MSI-H NEW |
Myeloid/lymphoid neoplasms for diagnosis, prognosis, and MRD support.
Aligned with ELN 2025 and enhanced CNV detection.
| Panel | Key Genes / Features |
|---|---|
| AML/MDS | FLT3, NPM1, DNMT3A, IDH1/2, RUNX1, ASXL1, TP53, TET2, SF3B1, SRSF2, CEBPA, BCOR, ETV6 NEW |
| MPN | JAK2, CALR, MPL, CSF3R |
| CLL | TP53, NOTCH1, SF3B1, BIRC3, MYD88; IGHV, BTK, PLCG2 NEW |
| ALL | IKZF1, CRLF2, JAK1/2, ABL1/2; fusions (ETV6-RUNX1, BCR-ABL1) |
| Lymphoma | MYD88, CD79B, EZH2, CARD11, BCL2/6 rearrangements |
ctDNA for actionable variants, tumor dynamics, MSI/TMB, and personalized MRD tracking.
| Panel | Key Features |
|---|---|
| Hotspot Panel | 20–100 genes, <1 ppm LOD NEW |
| Personalised MRD | Tumor-informed, RNA fusions (ALK, ROS1) NEW |
Actionable mutation-to-drug mapping based on approved and investigational targeted therapies. Designed for clinicians, researchers, and precision oncology programs.
| Mutation / Fusion | Cancer Type | Targeted Therapy | Mode of Action | Status |
|---|
⚠️ Therapy relevance depends on tumour type, line of therapy, and clinical guidelines (NCCN/ESMO).
We follow global standards (GDPR, HIPAA, ACMG, ICMR, NABL) to protect your rights, data, and well-being.
Questions? Email us at genetickundli@zohomail.in
Ready to begin? Share your details and our team will coordinate sample logistics and consent.